| Documentation | Build Status | DOI |
|---|---|---|
 |
||
 |
 |
The package is registered in the General registry so can be
installed with add. For example:
(@v1.6) pkg> add Phylo
Updating registry at `~/.julia/registries/General`
Updating git-repo `https://github.com/JuliaRegistries/General.git`
Resolving package versions...
Updating `~/.julia/environments/v1.6/Project.toml`
[aea672f4] + Phylo v0.4.18
Updating `~/.julia/environments/v1.6/Manifest.toml`
(@v1.6) pkg>The package is confirmed to work against Julia v1.6, the current LTS Julia v1.4 release and the latest release on Linux, macOS, and Windows. It is also tested against nightly.
Contributions are very welcome, as are feature requests and suggestions. Please open an issue if you encounter any problems or would just like to ask a question.
Phylo is a Julia package that provides
functionality for generating phylogenetic trees to feed into our
Diversity package to calculate phylogenetic
diversity. Phylo is currently in beta, but is probably still
missing much of the functionality that people may desire, so please
raise an issue if/when you find problems or missing
functionality - don't assume that I know!
Currently the package can be used to make trees manually, to generate
random trees using the framework from Distributions, and to read
newick and nexus format trees from files. It can also be used to
evolve continuous and discrete traits on the resultant phylogenies,
and plot all of this using Plots recipes. Finally, the trees and
traits are capable of handling Unitful units, so the branch lengths
can be time based, and traits that relate directly to physical units
(e.g. size) can be directly evolved.
For instance, to construct a sampler for 5 tip non-ultrametric trees, and then generate one or two random tree of that type (the examples below are from the dev branch, but work similarly on the current release):
julia> using Phylo
julia> nu = Nonultrametric(5);
julia> tree = rand(nu)
RootedTree with 5 tips, 9 nodes and 8 branches.
Leaf names are tip 2, tip 3, tip 1, tip 4 and tip 5
julia> trees = rand(nu, ["Tree 1", "Tree 2"])
TreeSet with 2 trees, each with 5 tips.
Tree names are Tree 2 and Tree 1
Tree 1: RootedTree with 5 tips, 9 nodes and 8 branches.
Leaf names are tip 5, tip 4, tip 2, tip 1 and tip 3
Tree 2: RootedTree with 5 tips, 9 nodes and 8 branches.
Leaf names are tip 3, tip 1, tip 2, tip 4 and tip 5The code also provides iterators, and filtered iterators over the branches, nodes, branchnames and nodenames of a tree, though this may soon be superseded by a simpler strategy.
julia> traversal(tree, inorder)
9-element Vector{LinkNode{OneRoot, String, Dict{String, Any}, LinkBranch{OneRoot, String, Dict{String, Any}, Float64}}}:
LinkNode tip 2, a tip of the tree with an incoming connection (branch 11).
LinkNode Node 7, an internal node with 1 inbound and 2 outbound connections (branches 13 and 11, 12)
LinkNode tip 3, a tip of the tree with an incoming connection (branch 12).
LinkNode Node 8, an internal node with 1 inbound and 2 outbound connections (branches 15 and 13, 14)
LinkNode tip 1, a tip of the tree with an incoming connection (branch 9).
LinkNode Node 6, an internal node with 1 inbound and 2 outbound connections (branches 14 and 9, 10)
LinkNode tip 4, a tip of the tree with an incoming connection (branch 10).
LinkNode Node 9, a root node with 2 outbound connections (branches 15, 16)
LinkNode tip 5, a tip of the tree with an incoming connection (branch 16).
julia> getnodename.(tree, traversal(tree, preorder))
9-element Vector{String}:
"Node 9"
"Node 8"
"Node 7"
"tip 2"
"tip 3"
"Node 6"
"tip 1"
"tip 4"
"tip 5"
julia> collect(nodenamefilter(isleaf, tree))
5-element Vector{String}:
"tip 1"
"tip 2"
"tip 3"
"tip 4"
"tip 5"The current main purpose of this package is to provide a framework for phylogenetics to use in our Diversity package, and they will both be adapted as appropriate until both are functioning as required (though they are currently working together reasonably successfully).
It can also read newick trees either from strings or files:
julia> using Phylo
julia> simpletree = parsenewick("((,Tip:1.0)Internal,)Root;")
RootedTree with 3 tips, 5 nodes and 4 branches.
Leaf names are Tip, Node 1 and Node 4
julia> getbranchname.(simpletree, getbranches(simpletree))
4-element Vector{Int64}:
1
2
3
4
julia> tree = open(parsenewick, Phylo.path("H1N1.newick"))
RootedTree with 507 tips, 1013 nodes and 1012 branches.
Leaf names are 227, 294, 295, 110, 390, ... [501 omitted] ... and 418And it can read nexus trees from files too:
julia> tree = open(parsenewick, Phylo.path("H1N1.newick"))
RootedTree with 507 tips, 1013 nodes and 1012 branches.
Leaf names are 227, 294, 295, 110, 390, ... [501 omitted] ... and 418
julia> ts = open(parsenexus, Phylo.path("H1N1.trees"))
[ Info: Created a tree called "TREE1"
[ Info: Created a tree called "TREE2"
TreeSet with 2 trees, each with 507 tips.
Tree names are TREE2 and TREE1
TREE1: RootedTree with 507 tips, 1013 nodes and 1012 branches.
Leaf names are H1N1_A_BRAZIL_11_1978, H1N1_A_TAHITI_8_1998, H1N1_A_TAIWAN_1_1986, H1N1_A_BAYERN_7_1995, H1N1_A_ENGLAND_45_1998, ... [501 omitted] ... and H1N1_A_PUERTORICO_8_1934
TREE2: RootedTree with 507 tips, 1013 nodes and 1012 branches.
Leaf names are H1N1_A_BRAZIL_11_1978, H1N1_A_TAHITI_8_1998, H1N1_A_TAIWAN_1_1986, H1N1_A_BAYERN_7_1995, H1N1_A_ENGLAND_45_1998, ... [501 omitted] ... and H1N1_A_PUERTORICO_8_1934
julia> ts["TREE1"]
RootedTree with 507 tips, 1013 nodes and 1012 branches.
Leaf names are H1N1_A_BRAZIL_11_1978, H1N1_A_TAHITI_8_1998, H1N1_A_TAIWAN_1_1986, H1N1_A_BAYERN_7_1995, H1N1_A_ENGLAND_45_1998, ... [501 omitted] ... and H1N1_A_PUERTORICO_8_1934
julia> gettreeinfo(ts)
Dict{String, Dict{String, Any}} with 2 entries:
"TREE2" => Dict("lnP"=>-1.0)
"TREE1" => Dict("lnP"=>1.0)
julia> gettreeinfo(ts, "TREE1")
Dict{String, Any} with 1 entry:
"lnP" => 1.0We so far only support calculating a few metrics on trees, but will gradually be added. Open an issue with a request!
julia> species = getleaves(tree)[[2, 5, 8, 12, 22]]; # take 5 tips from the phylogeny (or use names)
julia> mrca(tree, species) # Identify the MRCA (Most Recent Common Ancestor)
LinkNode Node 65, an internal node with 1 inbound and 2 outbound connections (branches 999 and 61, 62)And while we wait for me (or kind contributors!) to fill out
the other extensive functionality that many phylogenetics packages
have in other languages, the other important feature that it offers is
a fully(?)-functional interface to R, allowing any existing R library
functions to be carried out on julia trees, and trees to be read from
disk and written using R helper functions. Naturally the medium-term
plan is to fill in as many of these gaps as possible in Julia, so the R interface does not make RCall a dependency of the package (we use the
Requires package to avoid dependencies). Instead, if you want to use
the R interface you just need to use both packages:
julia> using Phylo
julia> using RCall
Creating Phylo RCall interface...
R> library(ape)You can then translate back and forth using rcopy on
R phylo objects, and RObject constructors on julia NamedTree
types to keep them in Julia or @rput to move the object into R:
julia> rt = rcall(:rtree, 10)
RCall.RObject{RCall.VecSxp}
Phylogenetic tree with 10 tips and 9 internal nodes.
Tip labels:
t10, t8, t1, t2, t6, t5, ...
Rooted; includes branch lengths.
julia> jt = rcopy(NamedTree, rt)
NamedTree with 10 tips, 19 nodes and 18 branches.
Leaf names are t8, t3, t7, t9, t6, ... [4 omitted] ... and t1
julia> rjt = RObject(jt); # manually translate it back to R
R> if (all.equal($rjt, $rt)) "no damage in translation"
[1] "no damage in translation"
julia> @rput rt; # Or use macros to pass R object back to R
julia> @rput jt; # And automatically translate jt back to R
R> jt
Phylogenetic tree with 10 tips and 9 internal nodes.
Tip labels:
t10, t8, t1, t2, t6, t5, ...
Rooted; includes branch lengths.
R> if (all.equal(rt, jt)) "no damage in translation"
[1] "no damage in translation"For the time being the code will only work with rooted trees with named tips and branch lengths. If there's demand for other types of trees, I'll look into it.
As far as traits are concerned, these can be continuous pr discrete. First a continuous trait:
julia> using Phylo, Plots, DataFrames, Random
julia> tree = rand(Nonultrametric(100)) # Defaults to mean tree depth of 1.0
RootedTree with 100 tips, 199 nodes and 198 branches.
Leaf names are tip 41, tip 7, tip 37, tip 81, tip 88, ... [94 omitted] ... and tip 89
julia> rand!(BrownianTrait(tree, "Trait"), tree) # Defaults to starting at 0.0, variance 1.0
RootedTree with 100 tips, 199 nodes and 198 branches.
Leaf names are tip 41, tip 7, tip 37, tip 81, tip 88, ... [94 omitted] ... and tip 89
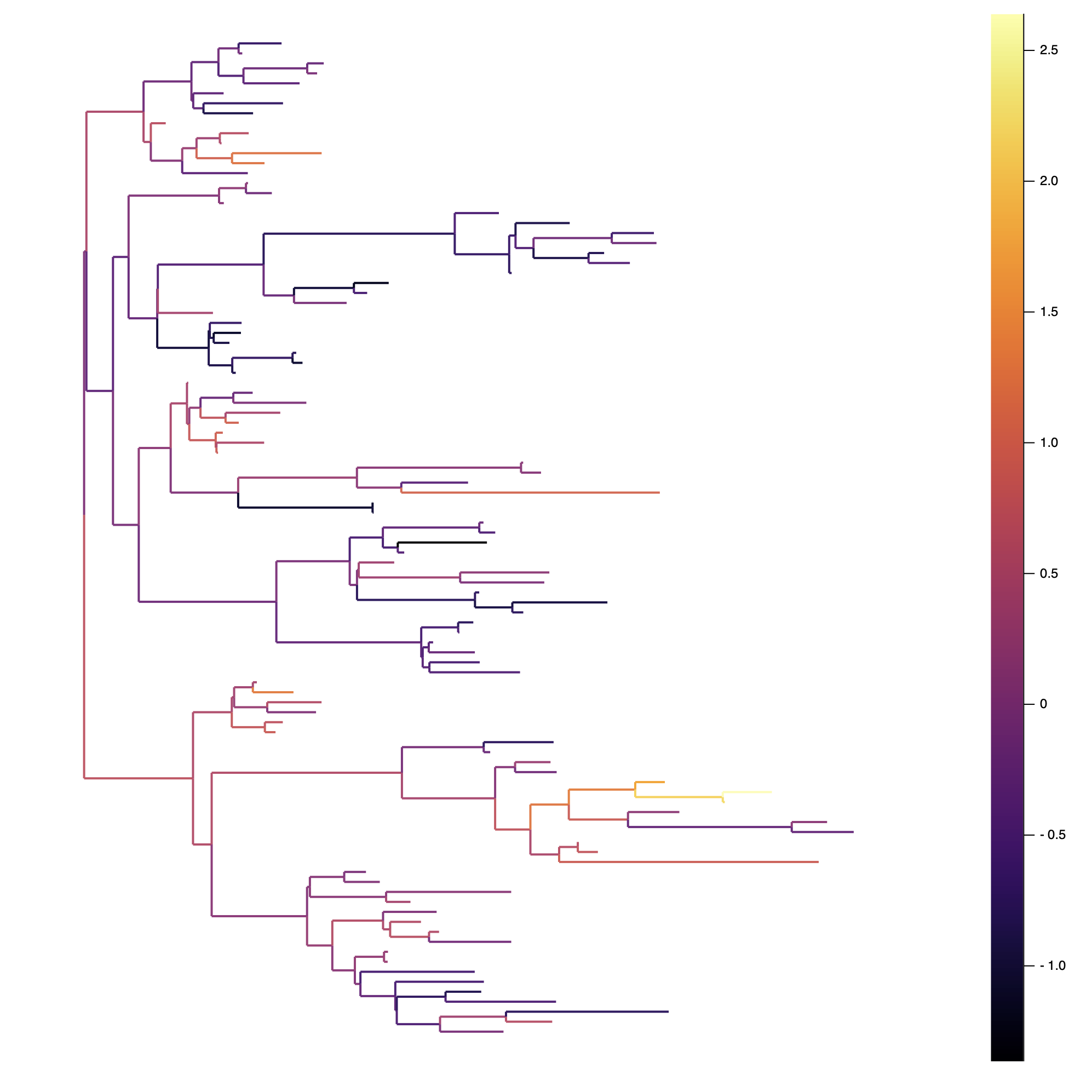
julia> plot(tree, line_z = "Trait", lw = 2)
julia> d = DataFrame(nodename=getnodename.(tree, traversal(tree, preorder)), trait=getnodedata.(tree, traversal(tree, preorder), "Trait"))
199×2 DataFrame
│ Row │ nodename │ trait │
│ │ String │ Float64 │
├─────┼──────────┼────────────┤
│ 1 │ Node 199 │ 0.0 │
│ 2 │ Node 198 │ 0.334372 │
│ 3 │ Node 163 │ 0.734348 │
│ 4 │ Node 138 │ 0.588014 │
│ 5 │ tip 41 │ 0.749697 │
⋮
│ 195 │ tip 64 │ -0.791095 │
│ 196 │ tip 75 │ -0.4173 │
│ 197 │ tip 82 │ -0.305623 │
│ 198 │ tip 71 │ -0.868774 │
│ 199 │ tip 89 │ -0.30126 │Then a discrete trait:
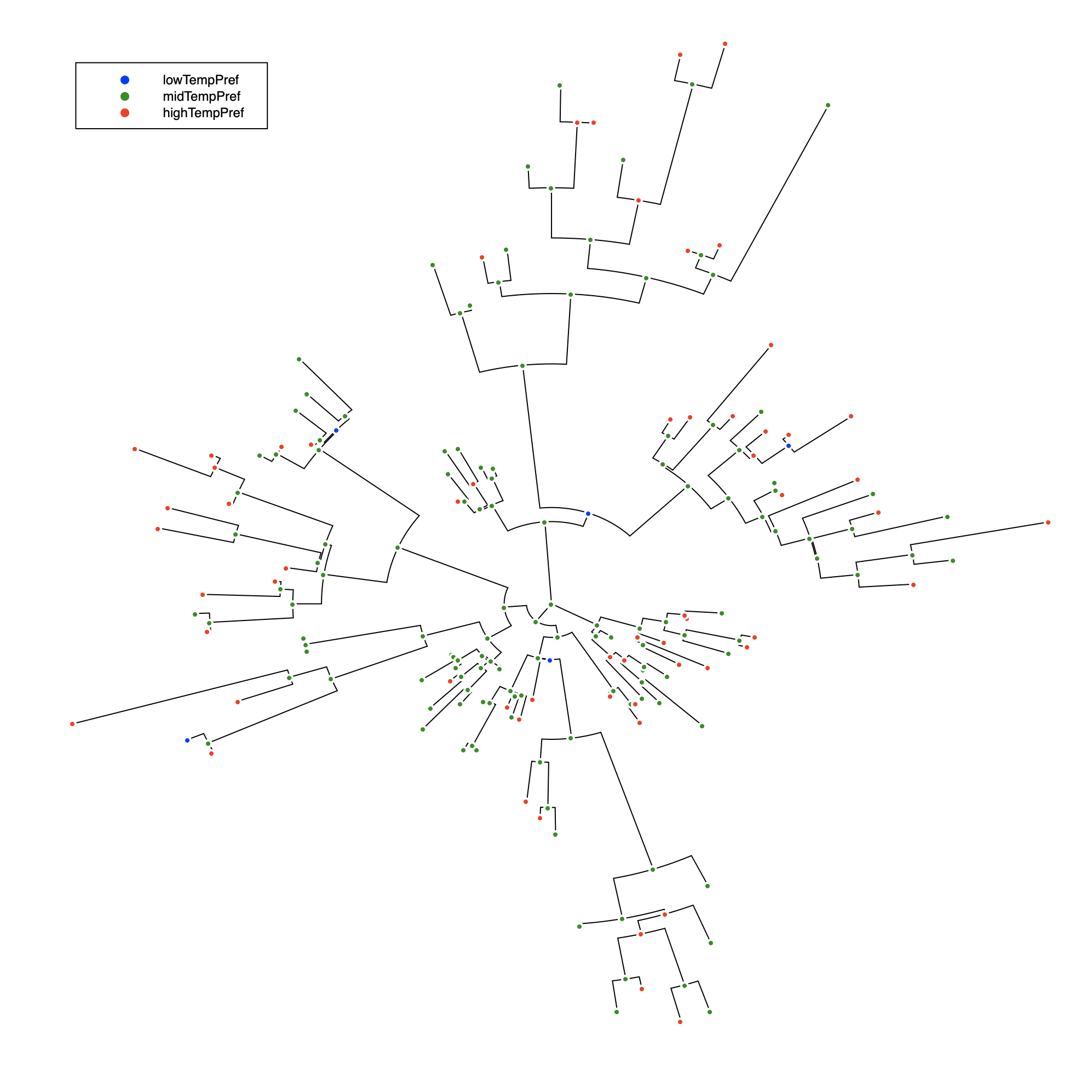
julia> @enum TemperatureTrait lowTempPref midTempPref highTempPref
julia> rand!(SymmetricDiscreteTrait(tree, TemperatureTrait, 0.4), tree);
julia> plot(tree, marker_group = "TemperatureTrait", legend = :topleft,
msc = :white, treetype = :fan, c = [:red :blue :green])
julia> d = DataFrame(nodename=getnodename.(tree, traversal(tree, preorder)), trait=getnodedata.(tree, traversal(tree, preorder), "TemperatureTrait"))
199×2 DataFrame
│ Row │ nodename │ trait │
│ │ String │ TemperatureTrait │
├─────┼──────────┼────────────────────────────────────┤
│ 1 │ Node 199 │ highTempPref::TemperatureTrait = 2 │
│ 2 │ Node 198 │ lowTempPref::TemperatureTrait = 0 │
│ 3 │ Node 163 │ lowTempPref::TemperatureTrait = 0 │
│ 4 │ Node 138 │ lowTempPref::TemperatureTrait = 0 │
│ 5 │ tip 41 │ lowTempPref::TemperatureTrait = 0 │
⋮
│ 195 │ tip 64 │ midTempPref::TemperatureTrait = 1 │
│ 196 │ tip 75 │ highTempPref::TemperatureTrait = 2 │
│ 197 │ tip 82 │ highTempPref::TemperatureTrait = 2 │
│ 198 │ tip 71 │ highTempPref::TemperatureTrait = 2 │
│ 199 │ tip 89 │ highTempPref::TemperatureTrait = 2 │