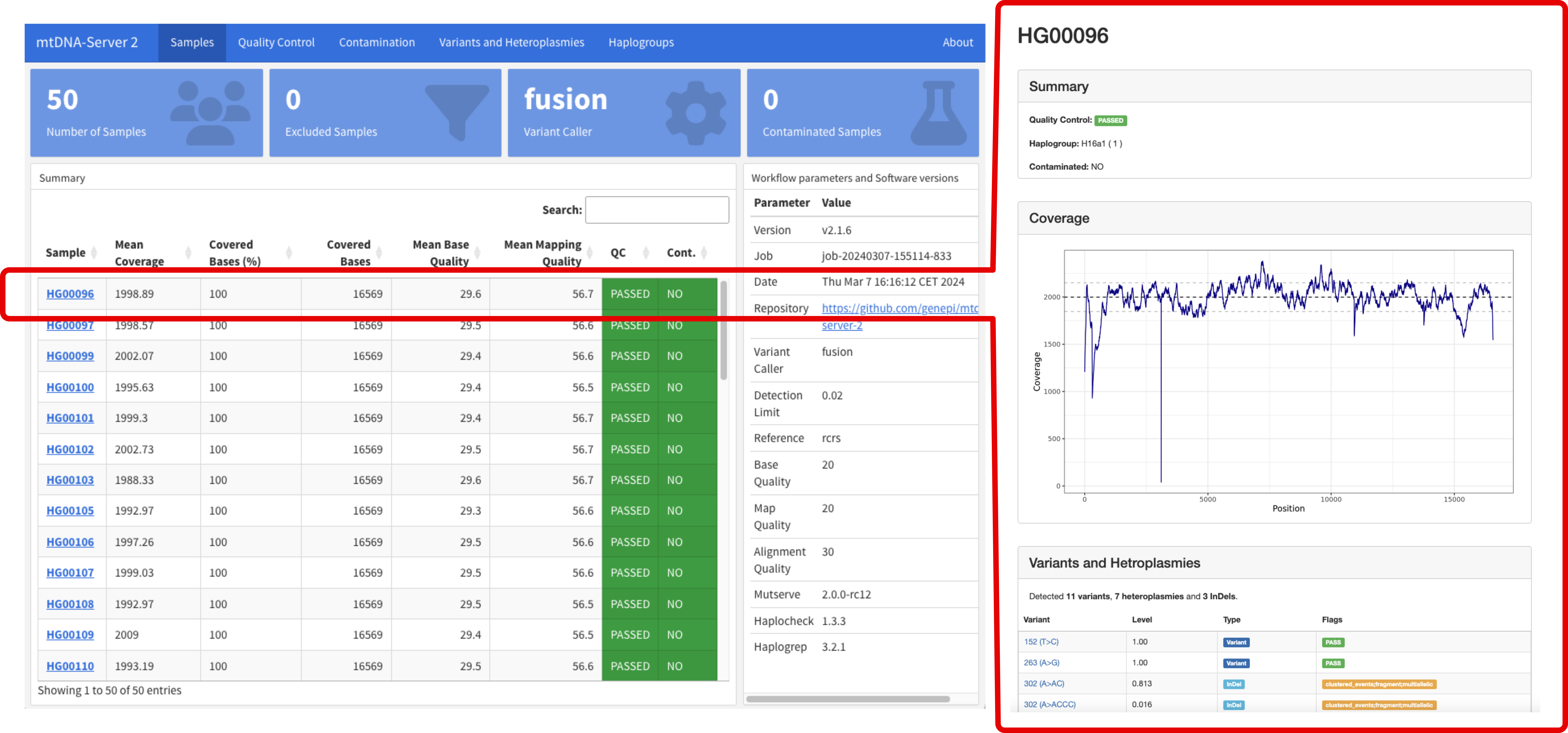
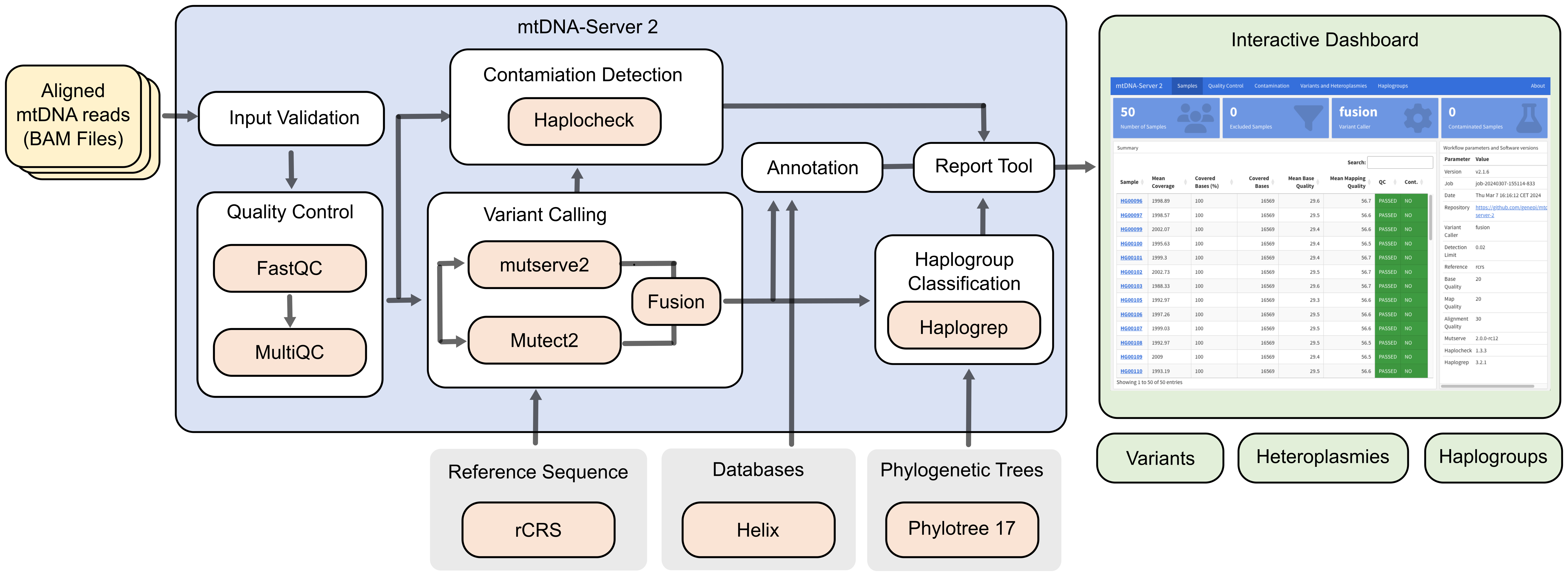
mtDNA-Server 2 is a Nextflow DSL2 pipeline to accurately detect heteroplasmic and homoplasmic variants in mitochondrial (mtDNA) genomes.
It supports different variant callers and is able to call insertions, deletions (INDELs) and single nucleotide variants (SNVs). Furthmore it includes several modules for (a) quality control and input validation (b) haplogroup classification and contamination detection, (c) minimal VAF estimation based on the coverage to minimize false positives and (d) an interactive analytics dashboard.
mtDNA-Server 2 is available as a graphical web-service or a Nextflow pipeline for local execution.
mtDNA-Server 2 is hosted as a free service on our mitoverse platform.
Documentation can be accessed here.
To run mtDNA-Server 2 locally, please execute the following steps.
-
Install Nextflow (>=22.10.4)
-
Run the pipeline on our test dataset and select either Docker, Singluarity or Slurm.
nextflow run genepi/mtdna-server-2 -r v2.1.9 -profile test,<docker,singularity,slurm>
To run mtDNA-Server 2 on your own data, create a config file and run the following command:
nextflow run genepi/mtdna-server-2 -r v2.1.9 -c <your-config-file> -profile docker
git clone https://github.com/genepi/mtdna-server-2
cd mtdna-server-2
docker build -t genepi/mtdna-server-2 . # don't ignore the dot
nextflow run main.nf -profile test,development
The following parameters can be set in the configuration file.
| Parameter | Default Value | Comment |
|---|---|---|
| project | null | Project name (required) |
| files | null | Input BAM files (required) |
| mode | fusion | Mode of operation (mutserve,mutect2,fusion) |
| detection_limit | 0.02 | Detection limit for heteroplasmic sites |
| coverage_estimation | on | Coverage estimation enabled |
| subsampling | off | Subsampling on/off |
| subsampling_coverage | 2000 | Subsampling coverage |
| mapQ | 20 | Mapping quality threshold |
| baseQ | 20 | Base quality threshold |
| alignQ | 30 | Alignment quality threshold |
| output | null | Specific Output folder |
Weissensteiner H*, Forer L*, Fuchsberger C, Schöpf B, Kloss-Brandstätter A, Specht G, Kronenberg F, Schönherr S: mtDNA-Server: next-generation sequencing data analysis of human mitochondrial DNA in the cloud. Nucleic Acids Res. 44:W64-9, 2016. PMID: 27084948
This software was developed at the Institute of Genetic Epidemiology, Medical University of Innsbruck
Sebastian Schoenherr (@seppinho)
Hansi Weissensteiner (@whansi)
MIT Licensed.